parallel blasts using python's pp module
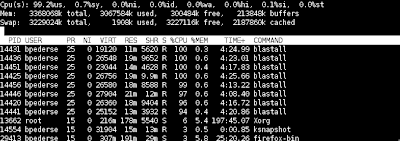
BLAST handles utilizes multiple cores for some scenarios using the -a flag. however, i often do full genome blasts -- blasting all chromosomes of one organism against all others. For the case of rice (O.s.) against itself, this is 124 jobs. there are simple tools in python to run a queued blast in which on my 8 core machine, each core will run 1 of those blasts, as a job finishes, the pp or parallel python module starts the next job, based on the number of cpus it has detected for your machine. the syntax for the script is: python pblast.py rice_rice_10kmers where rice_rice_10kmers is the section in a config/.ini file to get the parameters. My fasta directory looks like: $ ls /tmp/rice/fasta/ ricetenkmers_chr01.fasta ricetenkmers_chr04.fasta ricetenkmers_chr07.fasta ricetenkmers_chr10.fasta ricetenkmers.order ricetenkmers_chr02.fasta ricetenkmers_chr05.fasta ricetenkmers_chr08.fasta ricetenkmers_chr11.fasta ricetenkmers_chr03.fasta ricetenkmers_chr06.fasta ricetenkmers_chr...